Using Claude Code for EHR Informatics: Skills, Plugins, and MCP Servers, Part II
Part I was about context — orienting Claude to your work, your project. CLAUDE.md files, .claudeignore, reference tables, trusted queries. That foundation was important, but after a while, I noticed I was spending a lot of time re-explaining or re-orienting Claude: the same ICD query pitfalls, the same dsub machine type constraints, the same plotting conventions. CLAUDE.md files are a good way to bring some of this information into context, but context is also a finite resource. How do I efficiently bring in tools and expertise just when I need it?
Skills, plugins, hooks, and MCP servers are the way. This post walks through what I’ve built and what I’ve found useful from the community.
The series so far:
- Part I: Getting started — CLAUDE.md, .claudeignore, and why context matters
- Part II: Skills, plugins, hooks, and MCP servers ← you are here
- Part III: Building a cohort in All of Us with Claude Code
1. Skills: Stop Re-Explaining Yourself
A skill is a SKILL.md file — structured knowledge and workflows Claude can pull in when a situation calls for it. Unlike CLAUDE.md, which loads every session, skills load on demand. The tradeoff is intentional: CLAUDE.md gives Claude its baseline orientation, skills give it deep expertise in specific situations without loading everything at once.
Skills live in ~/.claude/skills/ for global use, or inside a project’s .claude/ directory for project-specific knowledge. The project-level location is where I put most of mine - it lets me titrate Claude’s abilities depending on whether I’m working on Scholia.fyi, the website I built to evaluate primary literature, or research-code, my repo for research tools.
Here’s a sample of what I’ve built recently.
My voice
bjw-voice-modeling is one of the rare global skills (~/.claude/skills/) that isn’t research-specific. Whether a drafted email or a notebook that I use to describe my work, I want it to sound like me. The skill includes four real writing samples across different contexts: an email to a colleague sharing null results, a methodological clarification correspondence, a speech I gave for a former senior resident, and a personal statement. Claude uses these to calibrate before writing anything on my behalf, orienting to my voice.
Anyone can build this, especially when you have Claude to help you. In addition to a bounty of helpful tools, Anthropic has a skill-creator skill that you can use to help compile your skill. For my voice modeling skill, I just provided the four writing samples and iterated with Claude to make this skill that I use almost daily.
Hopefully you’ll see this come up again and again - use Claude to make Claude better.
ICD queries
aou-icd-query builds on something I learned from a colleague. ICD code extraction in All of Us has a pitfall that isn’t obvious — one my labmate Tam Tran first identified when building PheTK: ICD-9 and ICD-10 both have codes beginning with “V,” but they mean entirely different things. Both share the same concept_code in the database, so a text join on condition_source_value returns both rows. (I mentioned this in Part I.)
Tam’s correct query has three stages: extract all ICD events from both condition_occurrence and observation (across both source value and source concept ID columns), then resolve V-code vocabulary ambiguity by tracing through concept_relationship, then union the results. Every time I started a new cohort, I was explaining this from scratch — or hoping Claude would reconstruct it correctly.
Now it’s in a skill, including a note I care about:
From the skill: “Prefer using the full SQL rather than importing the function — the three-stage structure is intentionally visible so users understand the V-code resolution and dual-table logic rather than treating it as a black box.”
The goal is legibility, not abstraction. These scripts help me try to teach as well as analyze in my notebooks.
dsub infrastructure
aou-dsub-infrastructure is institutional memory. Distributed computing on All of Us runs through dsub, which submits jobs to Google Cloud Batch. dsub has a few constraints that are painful to rediscover.
When we migrated to Google Batch last summer, one was particularly painful: c4 machine types didn’t work. c4 requires hyperdisk-balanced boot disks, and dsub couldn’t set boot disk types. I learned this by submitting a job, waiting for it to fail, and debugging the error - exactly the kind of thing that belongs in a skill rather than in someone’s head.
The skill includes a constraint table (machine families, provider requirements, network flags, logging paths), the dsub_script() function pattern I use across SAIGE GWAS and METAL meta-analysis workflows, job monitoring utilities, and a machine type selection guide drawn from actual projects. When I or a trainee opens a new genomics notebook, this doesn’t need to be re-established.
Table schemas
aou-table-schema is an interesting case. In Part I, I described putting All of Us CDR data dictionaries in _reference/all_of_us_tables/ as flat tsv files. Claude could read them when needed — but it had no guidance about when they were relevant or which file to use.
Wrapping those same files in a skill keeps the information out of context when it’s not needed, and adds routing logic: when looking up table columns, load table_schemas.tsv; when estimating query size, load table_row_counts.tsv; when you need to filter by care setting, load visit_concepts.tsv. For the larger files, the skill instructs Claude to grep first, then load. This keeps token usage down, but still makes the data available even when I forget to direct Claude to the reference material.
The shift from “here are files” to “here is a skill with files” is subtle. It turns passive storage into active guidance, but it really draws from why skills are so powerful - Claude has access to all skill metadata every time, so it knows where to look and when.

Plotting conventions
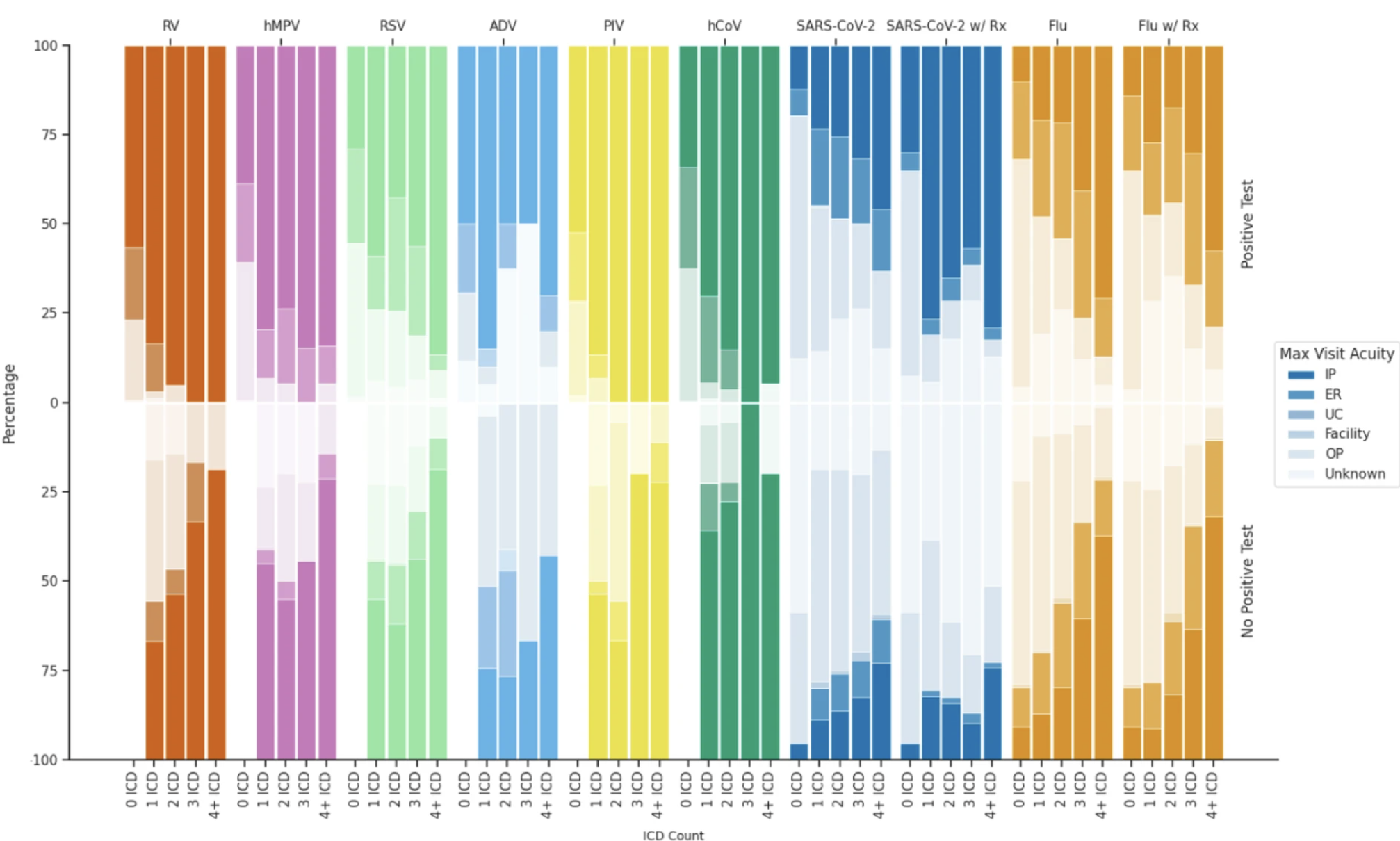
bjw-plotting is the smallest skill but saves real time. I care a lot about making figures that tell a story and look good. This skill carries forward my 11-color colorblind-friendly palette, the same seaborn whitegrid setup, consistent figure sizes by plot type. The All of Us count suppression rule also applies to plots: any annotation showing participant counts between 1 and 19 must display as "< 20". The skill encodes all of this, including the format_count() function, so I don’t establish it at the top of every notebook.
Here’s a taste from my last publication:
2. Plugins: Skills, Agents, Hooks, and More
Plugins are packaged distributions that “extend Claude Code with skills, agents, hooks, and MCP servers.” You install them once and they’re available across projects — no copying files between repos. They live in ~/.claude/plugins/.
The two sources I use:
example-skills
Anthropic ships a set of example skills as a plugin: document creation (docx, pptx, xlsx, pdf), frontend design, web artifacts, and more. These are genuinely useful for the non-code parts of research — formatted reports, slides, analysis dashboards. More importantly, this plugin is the reference implementation for how to write skills. If you want to build your own, reading through these teaches you the structure. The skill-creator skill (included here) guides you through the process.
The Life Sciences Plugin Marketplace
A more exciting set for researchers is the life sciences plugin marketplace. A few plugins worth knowing:
biorender: Integrates BioRender for scientific figure creation.
biorxiv: Access to preprints from bioRxiv and medRxiv. Useful when you want Claude to engage with recent work before it’s indexed in PubMed.
open-targets: This is my recent favorite. Open Targets aggregates SNP data from gnomad, GWAS associations, and QTL credible sets all in one place. For the question “what else is known about this locus?” — which I ask constantly — this replaces a round trip through multiple browser tabs with a structured answer in the same session where I’m already working.
nextflow-development: Guides users through nf-core pipelines with a structured checklist from data acquisition to output verification. I run genomics pipelines through dsub and PLINK2 on All of Us Researcher Workbench, so I haven’t used this one myself, but it targets exactly the bench scientist who has FASTQ files and needs to run a standardized pipeline.
3. Hooks: Guardrails For Your Code
CLAUDE.md instructions are soft constraints. Claude reads them, but in a long session with many steps, they can be forgotten or reasoned around. Hooks are hard constraints. They’re event-driven rules that intercept tool calls before they execute and warn or block based on patterns you define.
Blocking recursive GCS deletion
gsutil rm -r deletes everything at a GCS path, recursively. In an All of Us workspace, that could mean months of GWAS summary statistics, processed genotype files, or model outputs. There’s no recycle bin.
One hook I have intercepts any Bash command matching gsutil (-m )?rm -r and blocks it:
🛑 Recursive GCS deletion blocked!
You’re trying to recursively delete from Google Cloud Storage.
Why blocked: This can delete entire result directories.
Alternative: Move to a backup location instead — gsutil -m mv -r gs://bucket/old-results/ gs://bucket/trash/old-results-$(date +%Y%m%d)/
If I genuinely need to delete something, I can disable the rule temporarily, forcing an intentional extra safety step. Ten lines of YAML protecting months of work.
Creating hooks with hookify
I didn’t write that rule file by hand. hookify is a plugin that creates hooks from conversation — I ran /hookify, described what I wanted to prevent, and it generated the file. Rules take effect immediately, and if in practice I find the pattern to be too broad or too narrow, I edit the file directly. It’s worth installing early — once you see what hooks can do, you’ll want more of them.
4. MCP Servers: Real-Time External Connections
CLAUDE.md, skills, and reference files all provide static context — information that was true when you wrote it. (I of course should update my CLAUDE.md files, but this isn’t a strength of mine. Perhaps I need the right hook…) MCP servers connect Claude to external data. Claude can call external tools, get current results, and reason about them in the same context where you’re working.
Two I use somewhat regularly:
PubMed
The PubMed MCP gives Claude access to search_articles, get_full_text_article, find_related_articles, and get_article_metadata. While I do most of my recent literature search or review with Elicit or Scholia.fyi, this is a great resource. Instead of opening yet another tab, I use this to ask Claude to retrieve relevant papers and reason about methodology in the same session where I’m writing code.
context7
context7 retrieves current documentation for software libraries. When I’m working with a library I haven’t used recently — or one that’s changed since Claude’s training (#Polars once upon a time) — I ask Claude to pull the current docs before writing code. Library APIs shift, deprecated functions stick around in training data, and the difference between working code and code-that-looks-like-it-should-work is often one argument that changed between versions. context7 is a small habit that cuts time on avoidable debugging.
Building Your Own
If you want to go further:
Skills: Start with something specific — a query pattern you explain repeatedly, a constraint you keep rediscovering, a set of conventions you want consistent across projects. The skill-creator skill (in example-skills) guides you through the structure. Supporting reference files go in a references/ subdirectory alongside your SKILL.md.
Hooks: Run /hookify and describe what you want to prevent.
MCP servers: MCP servers are external processes that expose tools via a standardized protocol. The mcp-builder skill walks through creating one in Python or TypeScript.
What This Adds Up To
Part I was about giving Claude a map. Part II is about giving it tools. Together: Claude knows my project structure, my domain-specific query patterns, my coding conventions, my voice — and now it won’t delete my results directory, can search the literature without breaking context, and always uses the right palette for my figures.
That’s not replacing judgment. It’s scaffolding that takes the repetitive off my plate so judgment is what’s left. That’s the version of this I find worth using, and most of this can be found at my research-code repo. Hope they help!
In Part III, I’ll show what all of this looks like in practice: building a cohort from scratch in All of Us.
See you then.

Comments